 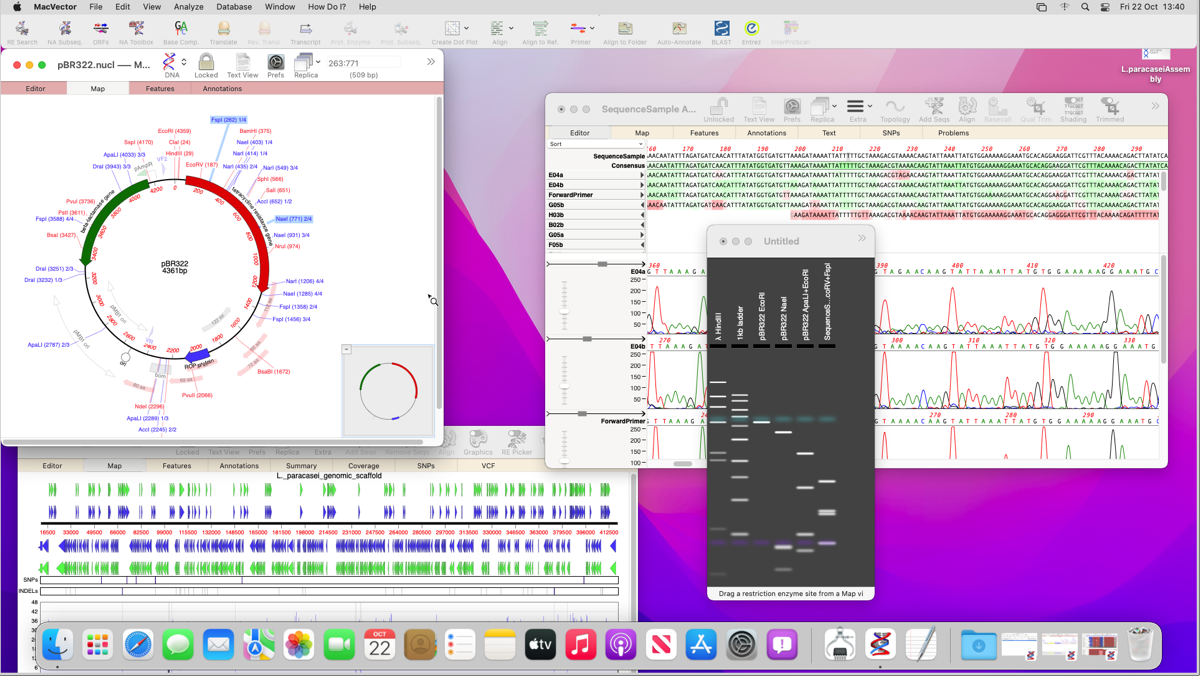
– If you suspect your primers may not be a perfect match then reduce SCORE THRESHOLD until your primer aligns. The resulting alignment will show your primers aligned against the template. You can switch to the Map view to show a graphical overview of all your primers and where they are located on the sequence. When to use: When you need to find many primers over a large sequenceīenefits: Easy to visualize a great quantity of primers against a template You can use the Editor view for a sequence level representation of the primer aligned against the template. Limitations: Your primer sequences need to be in a file already. This function is fairly easy to use and gives a large amount of detail about your primers. (i) Open your sequence and go to PRIMERS > Test PCR Primer Pairs For example secondary structure, what size the product will be and even the most ideal Tm for your PCR run. 
(ii) Copy and paste your two sequences in the two boxes. Note you will need to have pasted them into an external application. (iii) Click APPLY and see your primers detailed statistics. T-Coffee relies on your feeback.Primer 2: score 20, mismatches 0, lower strand 1592 to 1573 The 3' end of the primer binds within the product primer 1: score 20, mismatches 0, upper strand 1055 to 1074 Major product size: 538 bp Product details: (iv) Click OK to see the full statistics on the primers and product. T-Coffee is integrated in the MacVector sequence analys tool. 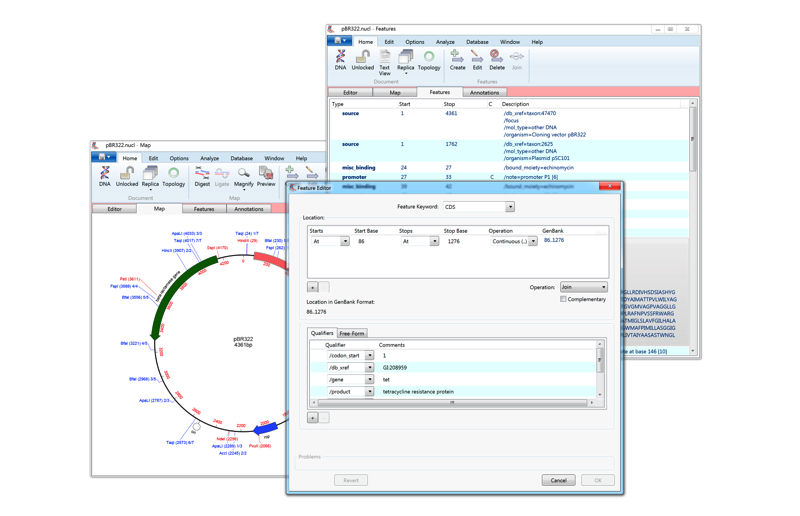
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
